from numpy.core import *
from numpy import r_, c_, diag
from numpy.random import permutation
from numpy.linalg import svd
from numpy.matrixlib import mat
import matplotlib.pyplot as plt
from scipy.spatial.distance import cdist
def findClosestCentroids(X: ndarray, centroids: ndarray):
K = size(centroids, 0)
idx = zeros((size(X, 0), 1))
idx = argmin(cdist(X, centroids), axis=1)
return idx
def computeCentroids(X: ndarray, idx: ndarray, K: int):
m, n = shape(X)
centroids = zeros((K, n))
for i in range(K):
centroids[i, :] = mean(X[nonzero(idx == i)[0], :], axis=0)
return centroids
def plotDataPoints(X, idx, K):
plt.scatter(X[:, 0], X[:, 1], c=idx)
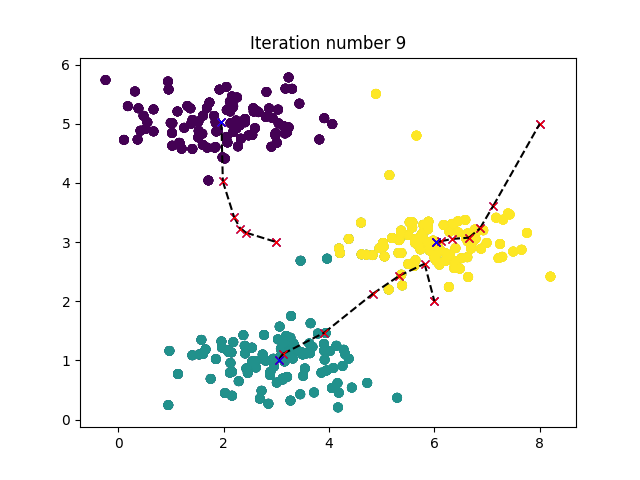
def plotProgresskMeans(X, centroids, previous, idx, K, i):
plotDataPoints(X, idx, K)
plt.plot(previous[:, 0], previous[:, 1], 'rx')
plt.plot(centroids[:, 0], centroids[:, 1], 'bx')
for j in range(size(centroids, 0)):
plt.plot(r_[centroids[j, 0], previous[j, 0]],
r_[centroids[j, 1], previous[j, 1]], 'k--')
plt.title('Iteration number {}'.format(i))
def runkMeans(X: ndarray, initial_centroids: ndarray, max_iters: int,
plot_progress=False):
if plot_progress:
plt.figure()
m, n = shape(X)
K = size(initial_centroids, 0)
centroids = initial_centroids.copy()
previous_centroids = centroids.copy()
idx = zeros((m, 1))
for i in range(max_iters):
print('K-Means iteration {}/{}...\n'.format(i, max_iters))
idx = findClosestCentroids(X, centroids)
if plot_progress:
plotProgresskMeans(X, centroids, previous_centroids, idx, K, i)
previous_centroids = centroids.copy()
centroids = computeCentroids(X, idx, K)
if plot_progress:
plt.show()
return centroids, idx
def kMeansInitCentroids(X: ndarray, K: int):
centroids = zeros((K, size(X, 1)))
randidx = permutation(size(X, 0))
centroids = X[randidx[:K, ], :]
return centroids
def featureNormalize(X: ndarray):
mu = mean(X, axis=0)
X_norm = X - mu
sigma = std(X_norm, axis=0, ddof=1)
X_norm /= sigma
return X_norm, mu, sigma
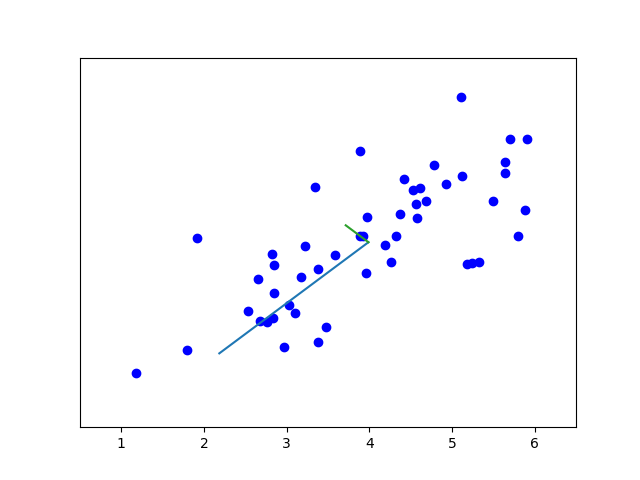
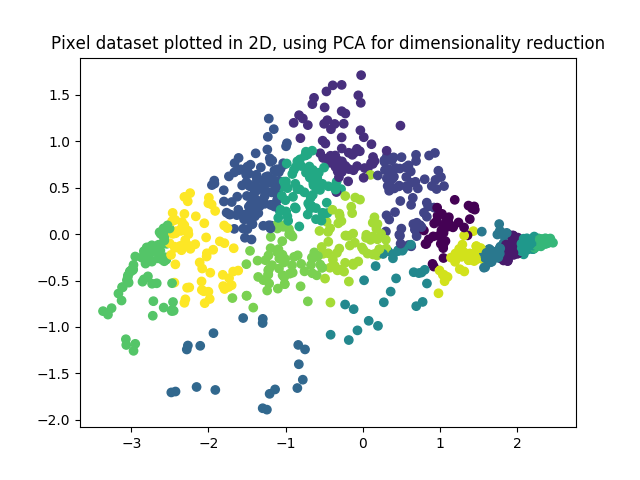
def pca(X: ndarray):
m, n = shape(X)
Sigma = X.T@ X/m
U, S, V = svd(Sigma)
return U, diag(S)
def drawline(p1, p2, *arg):
plt.plot(r_[p1[0], p2[0]], r_[p1[1], p2[1]], arg)
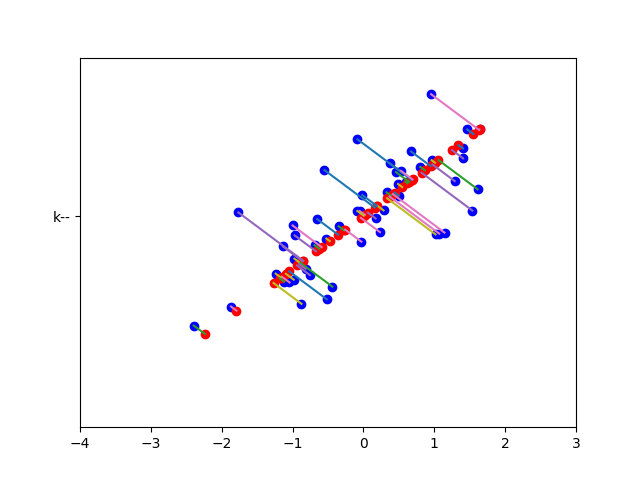
def projectData(X, U, K):
Z = zeros((size(X, 0), K))
U_reduce = U[:, : K]
Z = X @ U_reduce
return Z
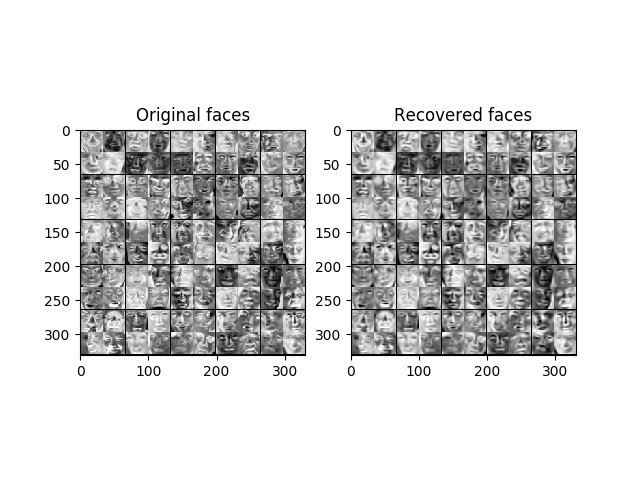
def recoverData(Z, U, K):
X_rec = zeros((size(Z, 0), size(U, 0)))
U_reduce = U[:, : K]
X_rec = Z @ U_reduce.T
return X_rec
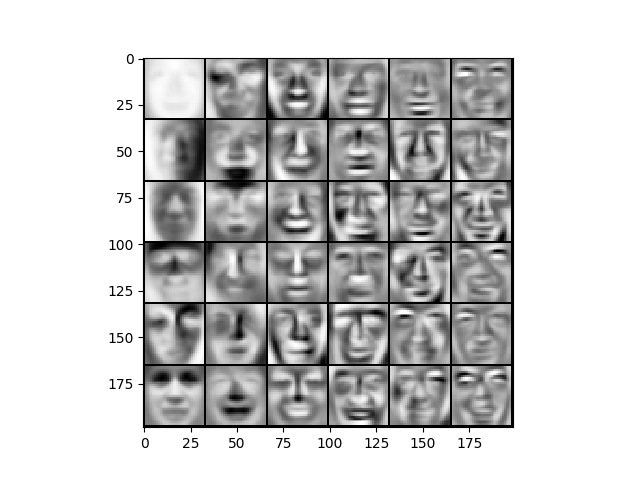
def displayData(X: ndarray, e_width=0):
if e_width == 0:
e_width = int(round(sqrt(X.shape[1])))
m, n = X.shape
e_height = int(n/e_width)
pad = 1
d_rows = int(floor(sqrt(m)))
d_cols = int(ceil(m / d_rows))
d_array = mat(
ones((pad+d_rows*(e_height+pad), pad + d_cols * (e_width+pad))))
curr_ex = 0
for j in range(d_rows):
for i in range(d_cols):
if curr_ex > m:
break
max_val = max(abs(X[curr_ex, :]))
d_array[pad+j*(e_height+pad) + 0:pad+j*(e_height+pad) + e_height,
pad+i*(e_width+pad)+0:pad+i*(e_width+pad) + e_width] = \
X[curr_ex, :].reshape(e_height, e_width)/max_val
curr_ex += 1
if curr_ex > m:
break
plt.imshow(d_array.T, cmap='Greys')
|